Spatial transcriptomics is one of the most exciting frontiers in biology today. This innovative technology allows us to map gene expression data directly onto tissue sections, giving us the ability to study biology at the intersection of genomics and tissue architecture. In other words, spatial transcriptomics allows scientists to understand not just which genes are active, but where they’re active within a tissue. Whether you’re exploring complex neurological networks or the intricate tumor microenvironment, analyzing spatial transcriptomics data can provide you with a wealth of insights that traditional sequencing methods simply can’t.
Before we get to the how, it’s important to first understand what spatial transcriptomics is. Simply put, spatial transcriptomics is a cutting-edge technique that allows researchers to measure gene expression while preserving the spatial context of tissues. This means that, instead of just identifying which genes are active in a sample, spatial transcriptomics gives us the ability to pinpoint exactly where those genes are active within a tissue section.
Traditional RNA-sequencing methods provide a snapshot of gene expression but don’t offer any information about where in the tissue that gene expression is happening. Spatial transcriptomics changes that by linking gene expression to spatial coordinates on tissue sections. This allows scientists to investigate complex tissue structures and understand how cells in a tissue communicate with one another based on their location.
Why is Spatial Transcriptomics Important?
- Spatial Context Matters: Tissue function isn’t just about which genes are expressed—it’s about where and how they interact within the tissue. For example, gene expression in the brain isn’t uniform; different regions of the brain have different functions, and understanding these patterns can be key to unlocking mysteries in neurobiology.
- Applications in Disease Research: In cancer research, for instance, spatial transcriptomics allows us to study the tumor microenvironment in great detail, observing how cancerous cells interact with surrounding healthy tissue. This is crucial for understanding how tumors grow and metastasize.
- Personalized Medicine: By analyzing tissue-specific gene expression, spatial transcriptomics can contribute to precision medicine by identifying individual tissue features that might influence treatment response.
A Brief History of Spatial Transcriptomics
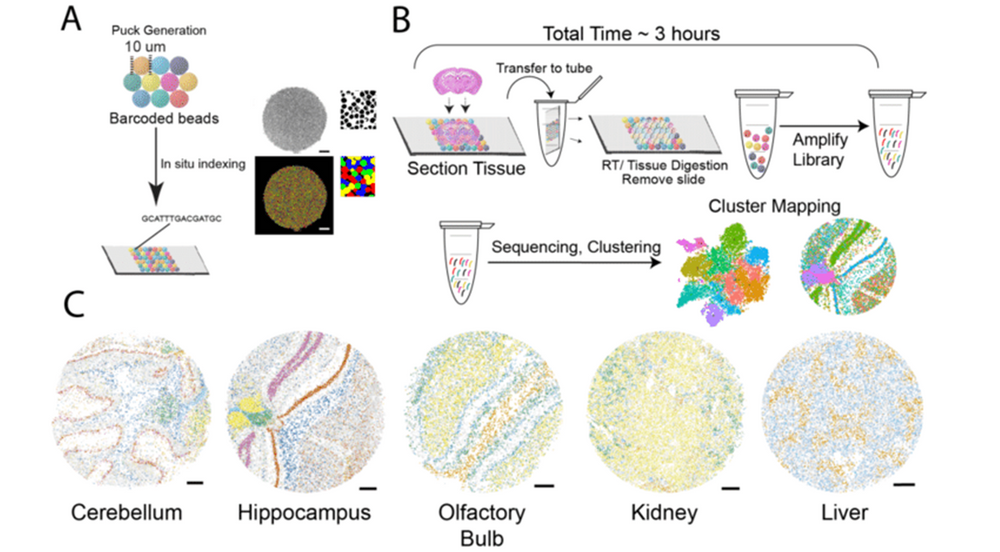
The concept of spatially resolved transcriptomics isn’t entirely new but has gained incredible momentum in recent years. The first papers on this topic emerged in the mid-2000s, but technological advances in sequencing, imaging, and microdissection have made spatial transcriptomics a more widely accessible tool. Today, platforms like 10x Genomics Visium, Slide-seq, and MERFISH offer high-resolution spatial mapping of gene expression.
What Does Spatial Transcriptomics Measure?
At its core, spatial transcriptomics measures gene expression across different spatial locations within tissue sections. But, there’s a little more to it than just reading RNA sequences!
Gene Expression in Spatial Context
Spatial transcriptomics takes a tissue section, typically a thin slice of biological tissue, and places it on a special slide where individual spatial spots are associated with a set of gene expression data. These “spots” can range from micron to millimeter sizes, depending on the resolution of the platform. Each spot contains RNA from the tissue located at that spot, and that RNA is sequenced to identify which genes are being expressed at that particular point.
Tissue Architecture
One of the unique aspects of spatial transcriptomics is that it doesn’t just provide a list of genes but gives us their spatial coordinates. This allows us to map gene expression data back to the tissue’s original 3D architecture, so we can observe the relationships between different cell types, tissues, and structures in the exact location they occupy.
Data Types in Spatial Transcriptomics
In practice, spatial transcriptomics generates two types of data:
- Gene Expression Data: A matrix of gene counts or expression levels for each spatial spot.
- Spatial Coordinates: Each spot in the tissue is assigned a set of spatial coordinates, essentially indicating where in the tissue that particular spot resides.
Common Platforms for Spatial Transcriptomics
Several technologies allow for the collection of spatial transcriptomics data, each with its own strengths and limitations:
- 10x Genomics Visium: This is currently one of the most popular platforms for spatial transcriptomics. It uses spatially barcoded tissue slides to capture gene expression in a spatially resolved manner, allowing for high-resolution analysis of tissues.
- Slide-seq: Another high-resolution technique that uses DNA barcodes to spatially tag RNA within tissue sections. This method is particularly noted for its ability to generate single-cell resolution data.
- MERFISH (Multiplexed Error-Robust Fluorescence In Situ Hybridization): A highly accurate method that uses fluorescence-based detection to measure gene expression with high spatial resolution. MERFISH allows for the simultaneous detection of hundreds to thousands of RNA species.
Why is Analyzing Spatial Transcriptomics Data So Important?
Analyzing spatial transcriptomics data provides key biological insights that traditional transcriptomics cannot. For example:
- Microenvironmental Influence: Gene expression can vary depending on the local microenvironment. For example, in tumors, the proximity to blood vessels or immune cells can influence gene expression patterns. Spatial transcriptomics allows you to study these interactions at a granular level.
- Spatial Heterogeneity: Tissue architecture can show varying gene expression profiles at different locations within the same tissue. Spatial transcriptomics uncovers this spatial heterogeneity, allowing us to study how certain genes might only be expressed in specific regions of the tissue, which could help us understand disease mechanisms or tissue development.
- Mapping Developmental Pathways: During development, tissues and organs undergo spatially regulated gene expression changes. By analyzing these patterns, we can understand how cells migrate and differentiate within the tissue architecture. This is especially important in developmental biology and neuroscience.
In short, spatial transcriptomics allows researchers to study biology as it occurs within the tissue context—something that was previously impossible with traditional methods. Now, we can take our analysis beyond “what genes are active?” and ask “where and how are those genes active?”
Laying the Groundwork for Data Analysis
Understanding how spatial transcriptomics works is key to analyzing the data that comes from these technologies. With its ability to map gene expression directly to tissue architecture, spatial transcriptomics opens up new doors for investigating complex biological systems.
But here’s the thing: The data can be overwhelming. There’s a lot of complexity involved in processing, analyzing, and interpreting spatial transcriptomics datasets. This is exactly why we need a systematic approach to analyze spatial transcriptomics data.
Preparing Your Data for Spatial Transcriptomics Analysis
Now that we’ve covered the basics of spatial transcriptomics and why it’s so important, let’s jump into how to get that data ready for analysis. This step is crucial because raw data—whether it’s from 10x Genomics Visium, Slide-seq, or any other platform—needs careful preprocessing to ensure the analysis goes smoothly.
Think of it like prepping ingredients before you cook: get everything in order, and your results will be much tastier. In the world of spatial transcriptomics, “tasty” means accurate, reproducible, and insightful biological data. So let’s start by looking at the key steps in preparing spatial transcriptomics data.
Preprocessing Raw Spatial Transcriptomics Data
When you first receive your spatial transcriptomics data, it may look like a mess of raw files. Don’t worry, this is normal. The first task is to clean up the data to make it ready for analysis. Here’s a breakdown of the key preprocessing steps:
- Data Quality Control (QC)
- The first thing you’ll want to do is check the quality of your data. Is there any noise? Are there any spots (data points) that don’t seem to match the expected patterns? Low-quality data can skew your results, so it’s important to filter it out before continuing.
- QC steps include:
- Gene filtering: If a gene is expressed in very few spots or has a very low count, it might be considered an outlier or background noise. These genes are usually removed in the preprocessing step.
- Spot filtering: Some spots might have little or no gene expression due to technical artifacts, such as empty spots or issues with the tissue slide. You can filter these out by setting a minimum threshold for gene expression or spot coverage.
Pro Tip: If you’re using software like Seurat or Scanpy, these platforms have built-in QC tools to help automate this process, making your life easier!
- Normalization
- Next up is normalization. Since tissues may differ in overall RNA content, it’s essential to normalize the data to ensure that observed gene expression patterns reflect real biological differences and not simply technical variability.
- There are different methods for normalization, such as:
- Library size normalization: Adjusting gene counts based on the total number of genes sequenced in each spot.
- Scaling: Adjusting the data so that each gene has a mean of 0 and a variance of 1 across all spots, ensuring that all genes are treated equally when analyzing expression.
Why this matters: Without normalization, differences in sequencing depth (how much RNA is captured from each spot) could falsely influence your results, leading to misleading conclusions about gene expression.
- Batch Effect Correction
- Batch effects can arise if samples were processed on different days or in different conditions, leading to systematic biases in your data. It’s crucial to identify and correct for these batch effects.
- Several tools exist to perform batch effect correction, such as Harmony or ComBat, which help ensure that your results reflect biological variation rather than technical artifacts.
Data Integration and Spatial Mapping
After you’ve completed the basic data quality checks, it’s time to link the data back to its spatial coordinates and tissue context. Here’s how you can approach it:
- Assigning Spatial Coordinates
- Each spot in your tissue section will have a set of spatial coordinates (X, Y) that corresponds to its position on the tissue slide. These coordinates are essential because they allow you to reconstruct the tissue architecture in relation to gene expression.
- Tools like Seurat or SpatialLIBS can help you easily integrate these spatial coordinates into your analysis framework. They enable you to visualize gene expression patterns in relation to specific regions of the tissue, which is key for understanding how genes are organized.
- Integration with Histological Data
- To gain a deeper understanding of the tissue’s structure, spatial transcriptomics data can be integrated with histological images (e.g., from Hematoxylin and Eosin (H&E) staining). These images give a visual representation of tissue types and structures, which can help guide your analysis.
- Some tools, like SpatialLIBS or SpatialDE, allow you to overlay gene expression maps on top of these histology images, providing a comprehensive view of how gene expression varies in different tissue areas.
Example: Imagine you’re studying brain tissue—by integrating spatial transcriptomics with neuroanatomical images, you could pinpoint how specific genes related to synaptic plasticity are expressed in different brain regions, helping to map functional areas of the brain.
Visualizing Spatial Data During Preprocessing
Visualization is an important step in preprocessing, as it allows you to visually inspect the data before moving on to the actual analysis. Here are some common visualization techniques to use:
- Spatial Heatmaps: These maps show the gene expression levels across the tissue, with darker or lighter shades indicating higher or lower expression, respectively. Spatial heatmaps allow you to get a quick look at how genes are distributed within tissue.
- t-SNE and UMAP Plots: While these are commonly used for dimensionality reduction in single-cell RNA-seq, they can also be applied to spatial transcriptomics data. These plots help to reduce the complexity of the data while retaining important spatial information, making it easier to identify patterns or clusters.Pro Tip: Tools like Scanpy, Seurat, and Matplotlib (for Python users) offer great visualization options for spatial data, and you can easily generate heatmaps, UMAPs, or even interactive plots to inspect your results.
Case Study: Preprocessing Data from 10x Genomics Visium
To make this more tangible, let’s look at an example. Imagine you have a dataset from 10x Genomics Visium, which provides high-resolution spatial transcriptomics data.
- Step 1: Perform QC filtering. You check the gene counts for each spot and remove spots with very few expressed genes.
- Step 2: Normalize the data. You apply library size normalization to account for differences in RNA content across the spots.
- Step 3: Integrate spatial coordinates with gene expression data, overlaying a H&E image to visualize tissue structure.
- Step 4: You create a spatial heatmap of gene expression in the hippocampus, which helps identify regions with higher expression of memory-related genes.
The result? A clean, normalized dataset where you can begin to perform deeper analyses like spatial clustering, differential gene expression, and ultimately gain insights into how different brain regions function in memory processes.
Preprocessing Sets the Stage for Success
Preprocessing is an essential first step in how to analyze spatial transcriptomics data. By carefully handling data quality control, normalization, and spatial integration, you ensure that your analysis will be robust, reproducible, and insightful. Think of it as setting the stage for a great performance—the data is now primed for deeper analysis!
Spatial Clustering and Pattern Recognition in Spatial Transcriptomics
Now that we’ve cleaned and preprocessed our spatial transcriptomics data, it’s time to move on to the next phase: clustering and pattern recognition. This is where the fun really begins! By analyzing how gene expression varies across the tissue, we can identify spatial domains, uncover biological patterns, and understand how different cell types interact within their environments.
Why Spatial Clustering Matters
Spatial transcriptomics gives us the unique opportunity to study gene expression in context—within the tissue’s natural architecture. But tissue is rarely homogenous. It’s made up of a variety of cell types, each with its own gene expression profile, and this variability can form spatially distinct regions, or clusters, within the tissue.
In this step, we aim to identify these regions of similar gene expression patterns. Some clusters might correspond to specific cell types, while others could reflect areas of tissue pathology, like a tumor or inflamed area. Identifying these patterns is critical for understanding how tissue architecture supports function, and can also help reveal new insights into disease progression.
How Does Spatial Clustering Work?
Spatial clustering in spatial transcriptomics is a bit like grouping people at a party based on their interests—except in this case, the “interests” are gene expression profiles, and the “party” is the tissue section. The idea is to group spots (or regions) with similar gene expression profiles, which can indicate a common function, developmental stage, or pathological state.
There are several clustering algorithms and methods commonly used for spatial transcriptomics analysis. Let’s explore the most common ones:
1. K-Means Clustering
- This is one of the simplest clustering algorithms, and it works by grouping spots into a predefined number (K) of clusters based on the similarity of their gene expression.
- Pros: Easy to implement and fast for small datasets.
- Cons: The choice of K (number of clusters) can be arbitrary and may not always reflect the underlying biological structure.
How it works: K-means assigns each data point (spot) to the nearest cluster center, iterating until the clusters are stable. While simple, K-means is often effective for identifying broad spatial patterns in tissue.
2. Graph-Based Clustering (Louvain Method)
- This method treats the spatial spots as nodes in a graph and connects them based on similarity in gene expression. The Louvain method is then used to identify communities (clusters) of closely connected spots.
- Pros: More flexible than K-means and works well with complex, irregular spatial structures.
- Cons: It can be computationally more intensive than K-means, especially for large datasets.
How it works: In the Louvain algorithm, spots that are closer in gene expression (and often in proximity in space) are grouped together, making it great for detecting subregions in tissues, such as areas of inflammation or distinct cell populations.
3. Hierarchical Clustering
- Hierarchical clustering doesn’t require you to set the number of clusters upfront. Instead, it builds a tree-like structure (dendrogram) by progressively grouping similar spots together, which can be cut at different levels to create different numbers of clusters.
- Pros: Doesn’t require the user to choose the number of clusters ahead of time, offering flexibility.
- Cons: Can be slower than other methods, especially with large datasets.
How it works: This method looks for hierarchical relationships between spots, creating clusters at different levels of granularity. It’s useful for discovering nested patterns in gene expression.
4. Spatially Constrained Clustering
- In spatial transcriptomics, spatial proximity matters. This means that spots that are close to one another (physically) may also have similar gene expression. Some algorithms, like SpatialDE and SpatialLIBS, incorporate spatial information directly into the clustering process, ensuring that spatial context is always considered when identifying clusters.
- Pros: More biologically accurate because it accounts for spatial dependencies between neighboring spots.
- Cons: May require more computational resources.
How it works: These algorithms prioritize clusters that are spatially cohesive, meaning they aim to identify regions of the tissue where gene expression is not only similar but also located close to one another in the tissue’s structure.
Visualizing Spatial Clusters
Once you’ve run your spatial clustering algorithm, the next step is to visualize the clusters. Visualization is key for interpreting the biological significance of these clusters and for communicating your findings. Here are some ways to effectively display spatial clusters:
- Spatial Heatmaps
- A spatial heatmap shows how gene expression varies across different spots, with color gradients representing the level of expression. You can overlay clusters on top of this heatmap to see how they correspond to different tissue regions.
- Clustered UMAP or t-SNE
- Although UMAP and t-SNE are typically used for dimensionality reduction in single-cell RNA-seq, they can also be applied to spatial transcriptomics. Once the data is clustered, these plots can show how the clusters are distributed across the tissue, providing a clearer picture of spatial gene expression patterns.
- Interactive Maps
- Interactive platforms like Seurat and Scanpy offer interactive maps, where users can hover over individual spots to view detailed gene expression data. This is great for exploration and deeper analysis.
Pro Tip: Using histology images in tandem with spatial clustering visualizations can enhance your understanding. For instance, in cancer research, you might visualize how tumor-associated clusters overlap with certain histological features, like tumor vasculature or immune infiltration.
Case Study: Identifying Tumor Microenvironment Clusters
Let’s say you’re analyzing a tumor sample using spatial transcriptomics, and you want to identify distinct clusters within the tumor and its surrounding microenvironment.
- Step 1: You apply a spatial clustering algorithm like the Louvain method to the gene expression data. The algorithm identifies several clusters within the tumor tissue, including regions of high immune cell activity and tumor proliferation.
- Step 2: You visualize these clusters on a spatial heatmap. This helps you see that certain areas of the tumor are highly active in genes related to angiogenesis (blood vessel formation), while other regions show high expression of immune checkpoint genes.
- Step 3: Finally, you overlay these clusters on top of a histology image, revealing that the angiogenic clusters are located near blood vessels, while the immune-active regions are found at the tumor’s periphery, suggesting that immune cells might be attempting to infiltrate the tumor.
The result? By clustering and visualizing gene expression patterns in this spatially resolved manner, you uncover new insights into how the tumor microenvironment is structured and how it interacts with immune cells.
Pattern Recognition: Detecting Gene Expression Hotspots
Spatial transcriptomics doesn’t just help with clustering; it also excels at identifying hotspots of gene expression. These hotspots represent areas in the tissue where certain genes are overexpressed or underexpressed in comparison to their surroundings. Detecting hotspots is critical for studying tissue development, disease progression, and local cellular interactions.
Some techniques for hotspot detection include:
- SpatialDE: This tool is designed to detect differential expression across space. It identifies genes that show spatially varying expression patterns and can be used to pinpoint hotspots or gradients of gene expression.
- Spatially Weighted Regression Models: These models are used to correlate gene expression with spatial position, allowing for the detection of expression patterns that change over distances in the tissue.
Why It’s Important: Hotspots of gene expression often correspond to key functional areas in the tissue, such as growth zones in developing organs, areas of active inflammation, or regions of cancerous transformation.
Clustering Reveals Tissue Function
Spatial clustering and pattern recognition are powerful tools that allow you to untangle the complexity of tissue organization. By identifying regions of similar gene expression, you can uncover how different cell types interact within their spatial context, revealing insights into normal tissue function as well as disease mechanisms. The ability to identify hotspots and clusters is what sets spatial transcriptomics apart from traditional transcriptomics, making it a game-changer for many areas of research.

Differential Gene Expression (DGE) Analysis in Spatial Context
With your data cleaned, normalized, and clustered, we’re now entering a critical phase of spatial transcriptomics analysis—differential gene expression (DGE) analysis. This step helps us determine which genes are expressed differentially between different regions of the tissue. In a typical transcriptomics experiment, DGE analysis tells you which genes are upregulated or downregulated in response to some condition. In spatial transcriptomics, however, we can do this with the added power of spatial context.
Why does this matter? Because gene expression isn’t always uniform across a tissue. Understanding where certain genes are over- or under-expressed within a tissue architecture can provide key insights into tissue function, disease mechanisms, and developmental pathways.
Why Spatial DGE Analysis is Important
In traditional transcriptomics, you often get an aggregate view of gene expression across a tissue sample. Spatial transcriptomics, however, lets us perform DGE analysis with spatial awareness. This allows us to:
- Identify Regional Variations: Different parts of the tissue may exhibit different gene expression profiles. For example, within a tumor, the core of the tumor might show upregulation of proliferation markers, while the edges of the tumor may express immune response genes.
- Study Tissue Architecture: Gene expression may vary depending on where the cells are located within the tissue. For instance, in a brain tissue sample, genes involved in synaptic transmission might be highly expressed in one region, while genes involved in neurogenesis might be more active in another.
- Uncover Disease Mechanisms: Understanding how gene expression differs across tissue regions can help reveal spatially regulated processes, like how tumors interact with their surrounding stroma or how developmental signaling gradients are set up.
In short, spatial DGE analysis gives you a more nuanced view of gene expression that traditional bulk RNA-seq simply can’t provide.
How to Perform Differential Gene Expression (DGE) in Spatial Transcriptomics
- Select the Regions for Comparison
- In spatial transcriptomics, the first step in DGE analysis is to decide which regions of the tissue you want to compare. This depends on the biological question you’re asking. For example:
- Comparing tumor vs. surrounding healthy tissue.
- Comparing different cell types within the same tissue (e.g., neurons vs. glial cells).
- Comparing different developmental stages within a tissue.
- In spatial transcriptomics, the first step in DGE analysis is to decide which regions of the tissue you want to compare. This depends on the biological question you’re asking. For example:
- Define Spatially Relevant Groups
- Once you’ve identified the regions of interest, group your spots into spatial regions. These could be based on clustering results or simply visual inspection of the tissue’s morphology (e.g., areas near blood vessels, immune cell infiltration zones, or tumor boundaries).
- Apply Differential Expression Tests
- After grouping your spots, the next step is to perform a differential expression test. Commonly used statistical tests for DGE analysis in spatial transcriptomics include:
- SpatialDE: A powerful tool specifically designed to detect genes whose expression varies spatially. It’s ideal for identifying genes that are differentially expressed between different tissue regions, accounting for spatial dependencies.
- SpatialLIBS: Another popular tool that handles spatially resolved differential expression. It accounts for both spatial correlation and variability in gene expression across tissue spots.
- DESeq2 or edgeR: Although these are traditionally used for bulk RNA-seq, they can be applied to spatial transcriptomics data with some modifications, though they don’t account for spatial dependencies as directly as the aforementioned tools.
- After grouping your spots, the next step is to perform a differential expression test. Commonly used statistical tests for DGE analysis in spatial transcriptomics include:
- Adjust for Spatial Autocorrelation
- Spatial autocorrelation refers to the phenomenon where spots located closer to each other are more likely to have similar gene expression levels. Since spatial transcriptomics data are inherently spatially correlated, it’s important to use statistical models that account for this correlation.
- Tools like SpatialDE or SPARK use spatial smoothing or neighbor-based models to ensure that the results of your DGE analysis aren’t biased by spatial proximity.
Visualizing Differential Gene Expression
Once the DGE analysis is complete, it’s time to visualize your results. Here are some effective ways to display spatially resolved differential expression:
- Spatial Heatmaps
- A spatial heatmap is a great way to show how the expression of a particular gene varies across tissue regions. These heatmaps often use color gradients to show gene expression levels, with hotter colors indicating higher expression.
- Volcano Plots
- Volcano plots are used to visualize the significance of differentially expressed genes. The x-axis represents the log fold change in expression (i.e., how much a gene’s expression increases or decreases between groups), while the y-axis represents the statistical significance. Highly significant genes with large fold changes are plotted toward the top-right or top-left corners, making it easy to spot important genes.
- t-SNE or UMAP Plots with Color Coding
- After performing DGE analysis, you can plot your data using dimensionality reduction techniques like t-SNE or UMAP. By color-coding the data points based on differential expression, you can see how genes are distributed spatially within the tissue and where significant expression differences occur.
- 3D Surface Maps
- For high-resolution datasets (like single-cell resolution), 3D surface plots can be created to show gene expression at different depths within the tissue, offering a more detailed spatial view of expression patterns.
Case Study: DGE Analysis of Tumor Tissue
Let’s put everything together with a case study. Suppose you’re studying breast cancer using spatial transcriptomics. You’ve obtained a high-resolution tissue section and run a DGE analysis comparing tumor regions with surrounding healthy tissue. Here’s how the process might unfold:
- Step 1: After preprocessing and spatial clustering, you identify two regions in the tumor tissue—one with high proliferation markers and another with immune cell infiltration.
- Step 2: You apply a spatial DGE tool like SpatialDE to identify which genes are upregulated in each region. You find that genes associated with angiogenesis (blood vessel growth) are upregulated in the proliferative region, while genes involved in immune checkpoint regulation are elevated in the immune-infiltrated area.
- Step 3: You visualize the results using spatial heatmaps, revealing that areas with high angiogenesis correspond to the center of the tumor, while immune-related genes are more active at the edges of the tumor.
- Step 4: By combining this DGE analysis with histology images, you see that the angiogenic region is closer to blood vessels, while the immune-related region corresponds to the tumor microenvironment’s periphery, where immune cells are likely attempting to penetrate.
This analysis gives you a clear, spatially resolved picture of how tumor regions are interacting with their microenvironment—information that could be used to develop targeted therapies.
DGE Analysis Unlocks the Power of Spatial Transcriptomics
Differential gene expression analysis in spatial transcriptomics adds a whole new layer of insight to your data by allowing you to assess gene expression with respect to the tissue’s spatial architecture. By identifying differentially expressed genes in different tissue regions, you can uncover important biological patterns related to tissue function, disease progression, and cellular interactions.
This step is essential for truly understanding how gene expression drives tissue development, health, and disease at the spatial level.